Bigdata – In English
Data from ONERBA’s networks are now freely available on line through dedicated APIs.
Human data are available in the BIGDATA API, available HERE and functionalities are described underneath.
For animal data, a second API, R-SHINY, has been developed by the Resapath network on the ANSES website as it was very complex to gather animal and human data in a single database.
BIGDATA APPLICATION – Resistance data from human infections
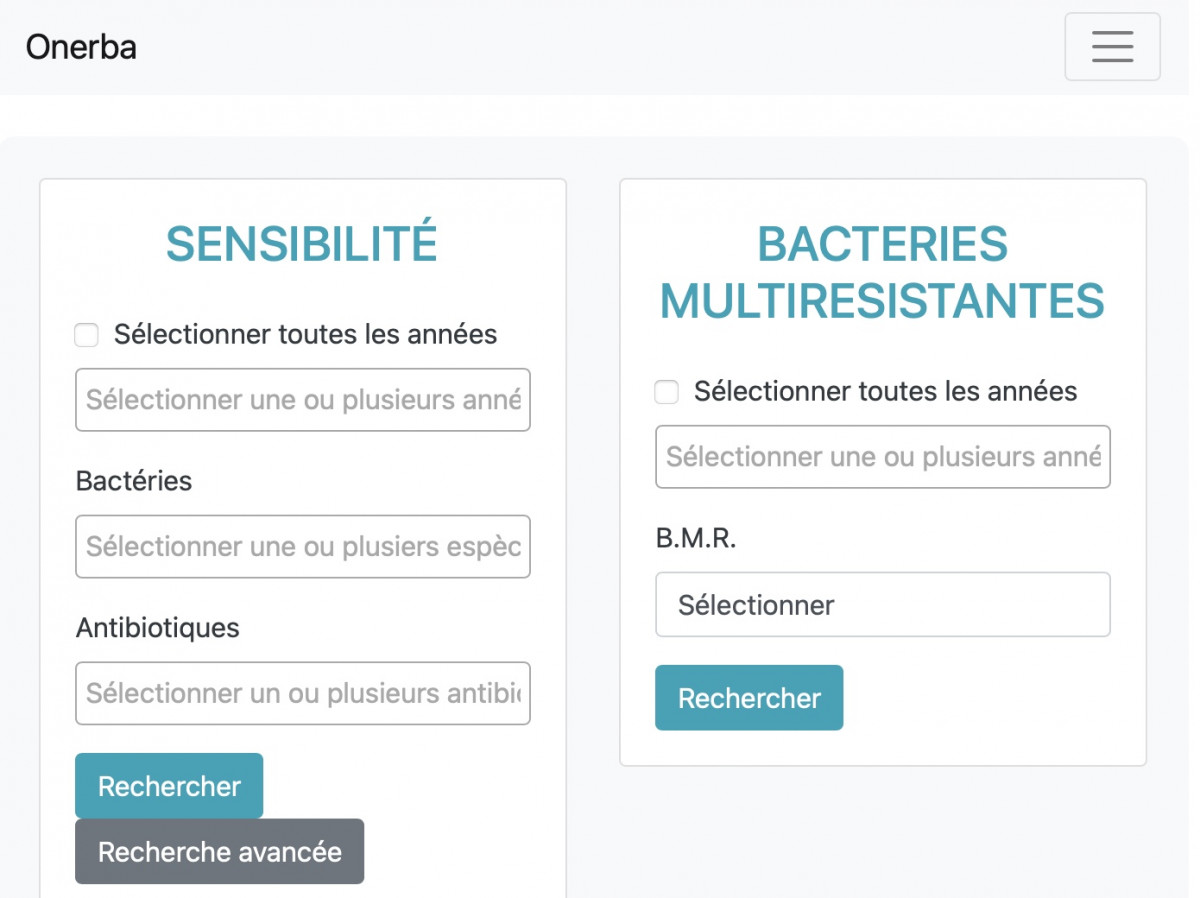
- The API allows finding susceptibility data for the main species of clinical interest in human pathology
- presented as global resistance
- and by using an advanced research, according to numerous characteristics (age groups, gender, type of clinical sample, hospital- or communitary-acquired …)
- Searching for multidrug resistant organisms (MDR) is also available.
- Finally, analysis of diameters or MICs distributions (not available cet) is available for selected couples [species/antibiotic].
- Whenever possible, time trends are presented when multiple years are selected.
Data are tabulated as % of susceptibility (excepted for MDR and MCIs or diameters distributions).
Tables and figures are downloadable and can be used for free providing acknowledging the source of data.
ATTENTION – data should be interpreted according to the methodology used by each network. Details are available on network pages.
OF NOTE
- Only couples [bacterial species/antibiotic] with ≥ 30 tests are considered for analysis.
- All data collected by the networks are not available in the database and we are working hard to complete BIGDATA.
Data collected by some networks will never be available in BIGDATA database:
- Either because the network did not grant access to its database
- or because the format cannot fit in the standardized format necessary for entering data in the API.
Each network is responsible for the collection, cleaning, and availability of its database.
Despite our efforts, BIGDATA may display some errors or bugs. We thank you to help us to improve the API by keeping us informed about your difficulties.